W#09: Hypothesis Testing, Classification and Regression with Decision Trees, Overfitting
Packages used here
Hypothesis testing
Large part of the content adapted from http://datasciencebox.org.
Organ donors
Background story: People providing an organ for donation sometimes seek the help of a special “medical consultant”. These consultants assist the patient in all aspects of the surgery, with the goal of reducing the possibility of complications during the medical procedure and recovery. Patients might choose a consultant based in part on the historical complication rate of the consultant’s clients.
Example track record: One consultant tried to attract patients by noting that the average complication rate for liver donor surgeries in the US is about 10%, but her clients have only had 3 complications in the 62 liver donor surgeries she has facilitated. She claims this is strong evidence that her work meaningfully contributes to reducing complications (and therefore she should be hired!).
Is this strong evidence for a good track record?
Data
Parameter vs. statistic
A parameter for a hypothesis test is the “true” value of interest. We typically estimate the parameter using a sample statistic as a point estimate.
\(q\): true rate of complication, here 0.1 (10% complication rate in US)
\(\hat{q}\): rate of complication in the sample = \(\frac{3}{62}\) = 0.048 (This is the point estimate.)
Correlation vs. causation
Is it possible to infer the consultant’s claim using the data?
No. The claim is: There is a causal connection. However, the data are observational. For example, maybe patients who can afford a medical consultant can afford better medical care, which can also lead to a lower complication rate (for example).
What is possible?
While it is not possible to assess the causal claim, it is still possible to test for an association using these data. For this question we ask, could the low complication rate of \(\hat{q}\) = 0.048 be due to chance?
Two claims
- Null hypothesis \(H_0\): “There is nothing going on”
Complication rate for this consultant is no different than the US average of 10%
- Alternative hypothesis \(H_1\): “There is something going on”
Complication rate for this consultant is lower than the US average of 10%
Understand hypothesis testing as a court trial
Null hypothesis, \(H_0\): Defendant is innocent
Alternative hypothesis, \(H_A\): Defendant is guilty
Present the evidence: Collect data
Judge the evidence: “Could these data plausibly have happened by chance if the null hypothesis were true?”
- Yes: Fail to reject \(H_0\)
- No: Reject \(H_0\)
Hypothesis testing framework
Start with a null hypothesis, \(H_0\), that represents the status quo
Set an alternative hypothesis, \(H_A\), that represents the research question, i.e. what we are testing for
Conduct a hypothesis test under the assumption that the null hypothesis is true and calculate a p-value.
Definition p-value: Probability of observed or more extreme outcome given that the null hypothesis is true.- if the test results suggest that the data do not provide convincing evidence for the alternative hypothesis, stick with the null hypothesis
- if they do, then reject the null hypothesis in favor of the alternative
Setting the hypotheses
Which of the following is the correct set of hypotheses for the claim that the consultant has lower complication rates?
\(H_0: q = 0.10\); \(H_A: q \ne 0.10\)
\(H_0: q = 0.10\); \(H_A: q > 0.10\)
\(H_0: q = 0.10\); \(H_A: q < 0.10\)
\(H_0: \hat{q} = 0.10\); \(H_A: \hat{q} \ne 0.10\)
\(H_0: \hat{q} = 0.10\); \(H_A: \hat{q} > 0.10\)
\(H_0: \hat{q} = 0.10\); \(H_A: \hat{q} < 0.10\)
- is correct.
Hypotheses are about the true rate of complication \(q\) not the observed ones \(\hat{q}\)
Simulation for Hypothesis Testing
Simulating the null distribution
Since \(H_0: q = 0.10\), we need to simulate a null distribution where the probability of success (complication) for each trial (patient) is 0.10.
How should we simulate the null distribution for this study using a bag of chips?
- How many chips? For example 10 which makes 10% choices possible
- How many colors? 2
- What should colors represent? “complication”, “no complication”
- How many draws? 62 as the data
- With replacement or without replacement? With replacement
When sampling from the null distribution, what would be the expected proportion of “complications”? 0.1
Simulation!
set.seed(1234) # The seed is set for the sake of stable presentation outcomes.
outcomes <- c("complication", "no complication")
sim1 <- sample(outcomes, size = 62, prob = c(0.1, 0.9), replace = TRUE)
sim1 [1] "no complication" "no complication" "no complication" "no complication"
[5] "no complication" "no complication" "no complication" "no complication"
[9] "no complication" "no complication" "no complication" "no complication"
[13] "no complication" "complication" "no complication" "no complication"
[17] "no complication" "no complication" "no complication" "no complication"
[21] "no complication" "no complication" "no complication" "no complication"
[25] "no complication" "no complication" "no complication" "complication"
[29] "no complication" "no complication" "no complication" "no complication"
[33] "no complication" "no complication" "no complication" "no complication"
[37] "no complication" "no complication" "complication" "no complication"
[41] "no complication" "no complication" "no complication" "no complication"
[45] "no complication" "no complication" "no complication" "no complication"
[49] "no complication" "no complication" "no complication" "no complication"
[53] "no complication" "no complication" "no complication" "no complication"
[57] "no complication" "no complication" "no complication" "no complication"
[61] "no complication" "no complication"[1] 0.0483871Oh OK, this was exactly the consultant’s rate. But maybe it was a rare event? Let us repeat it.
More simulation!
Automating with tidymodels1
set.seed(10)
null_dist <- organ_donor |>
specify(response = outcome, success = "complication") |>
hypothesize(null = "point",
p = c("complication" = 0.10, "no complication" = 0.90)) |>
generate(reps = 100, type = "draw") |>
calculate(stat = "prop")
null_distResponse: outcome (factor)
Null Hypothesis: point
# A tibble: 100 × 2
replicate stat
<int> <dbl>
1 1 0.0323
2 2 0.0645
3 3 0.0968
4 4 0.0161
5 5 0.161
6 6 0.0968
7 7 0.0645
8 8 0.129
9 9 0.161
10 10 0.0968
# ℹ 90 more rowsVisualizing the null distribution
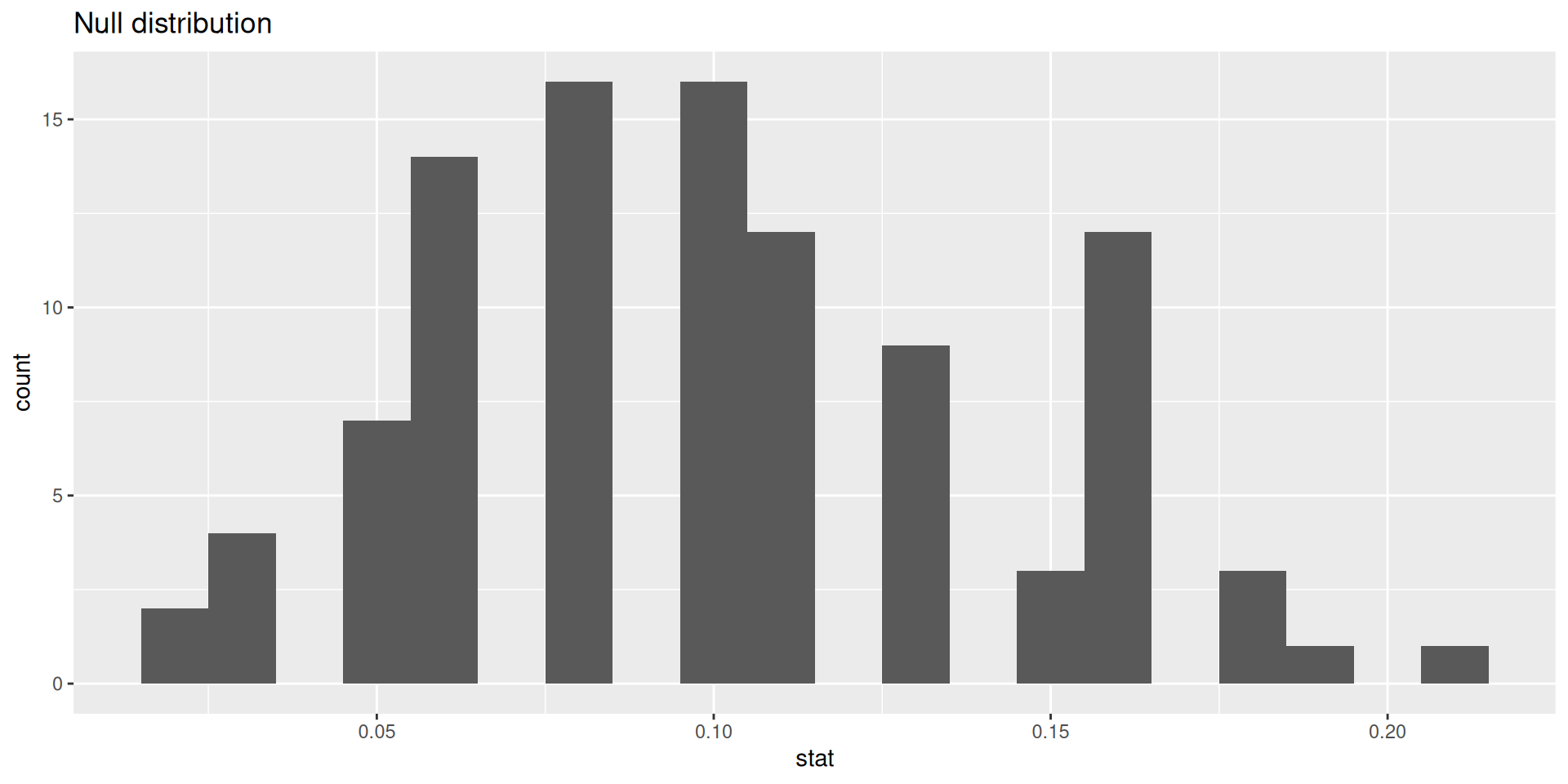
p-value
Calculating the p-value, visually
What is the p-value: How often was the simulated sample proportion at least as extreme as the observed sample proportion?1
Calculating the p-value, directly
This is the fraction of simulations where complications was equal or below 0.0483871.
Significance level
A significance level \(\alpha\) is a threshold we make up to make our judgment about the plausibility of the null hypothesis being true given the observed data.
We often use \(\alpha = 0.05 = 5\%\) as the cutoff for whether the p-value is low enough that the data are unlikely to have come from the null model.
If p-value < \(\alpha\), reject \(H_0\) in favor of \(H_A\): The data provide convincing evidence for the alternative hypothesis.
If p-value > \(\alpha\), fail to reject \(H_0\) in favor of \(H_A\): The data do not provide convincing evidence for the alternative hypothesis.
What is the conclusion of the hypothesis test?
Since the p-value is greater than the significance level, we fail to reject the null hypothesis. These data do not provide convincing evidence that this consultant incurs a lower complication rate than the 10% overall US complication rate.
100 simulations is not sufficient
- We simulate 15,000 times to get an accurate distribution.
null_dist <- organ_donor |>
specify(response = outcome, success = "complication") |>
hypothesize(null = "point",
p = c("complication" = 0.10, "no complication" = 0.90)) |>
generate(reps = 15000, type = "simulate") |>
calculate(stat = "prop")
ggplot(data = null_dist, mapping = aes(x = stat)) +
geom_histogram(binwidth = 0.01) +
geom_vline(xintercept = 3/62, color = "red")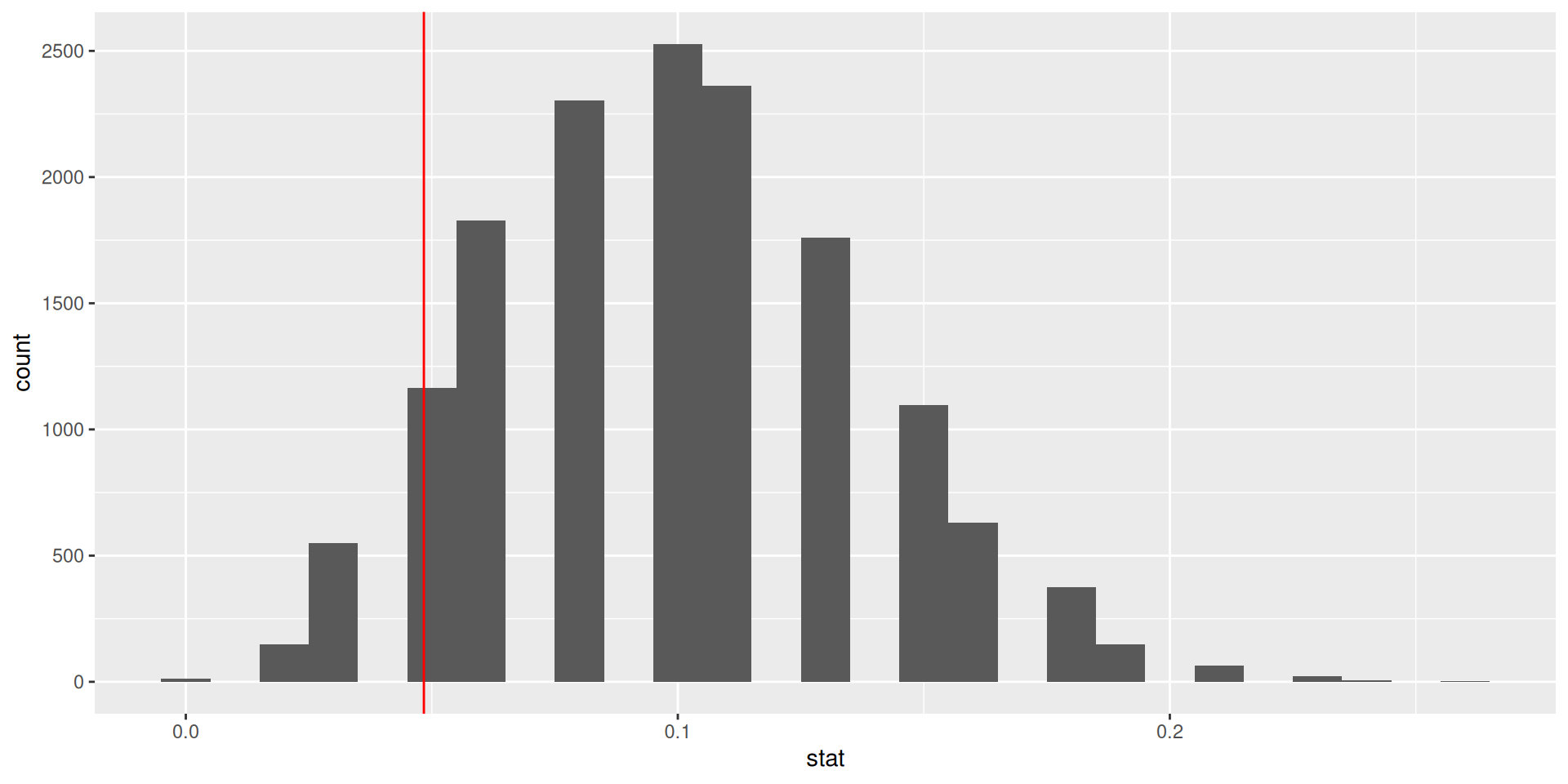
Our more robust p-value
For the null distribution with 15,000 simulations
# A tibble: 1 × 1
p_value
<dbl>
1 0.125Oh OK, our first p-value was much more borderline in favor of the alternative hypothesis.
p-value in model outputs
Model output for a linear model with palmer penguins.
# A tibble: 2 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 55.1 2.52 21.9 6.91e-67
2 bill_depth_mm -0.650 0.146 -4.46 1.12e- 5Model output for a logistic regression model with email from openintro
logistic_reg() |> set_engine("glm") |>
fit(spam ~ from + cc, data = email, family = "binomial") |> tidy()# A tibble: 3 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 13.6 309. 0.0439 0.965
2 from1 -15.8 309. -0.0513 0.959
3 cc 0.00423 0.0193 0.220 0.826What do the p-values mean? What is the null hypothesis?
Null-Hypothesis: There is no relationship between the predictor variable and the response variable, that means that the coefficient is equal to zero.
Smaller p-value \(\to\) more evidence for rejecting the hypothesis that there is no effect.
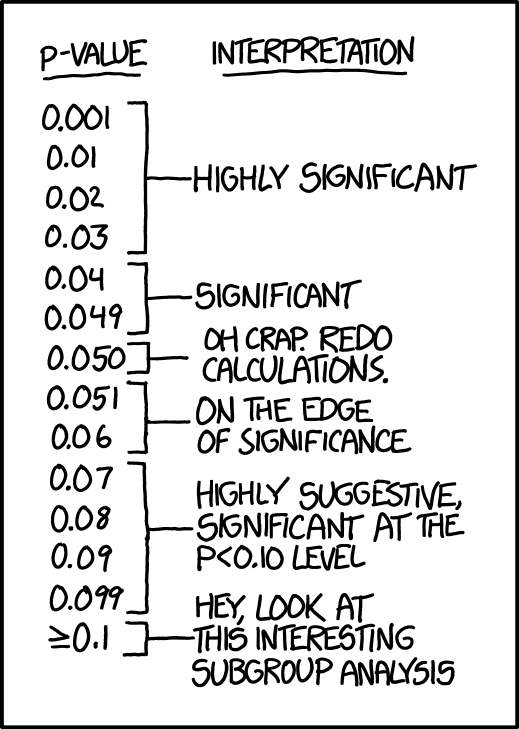
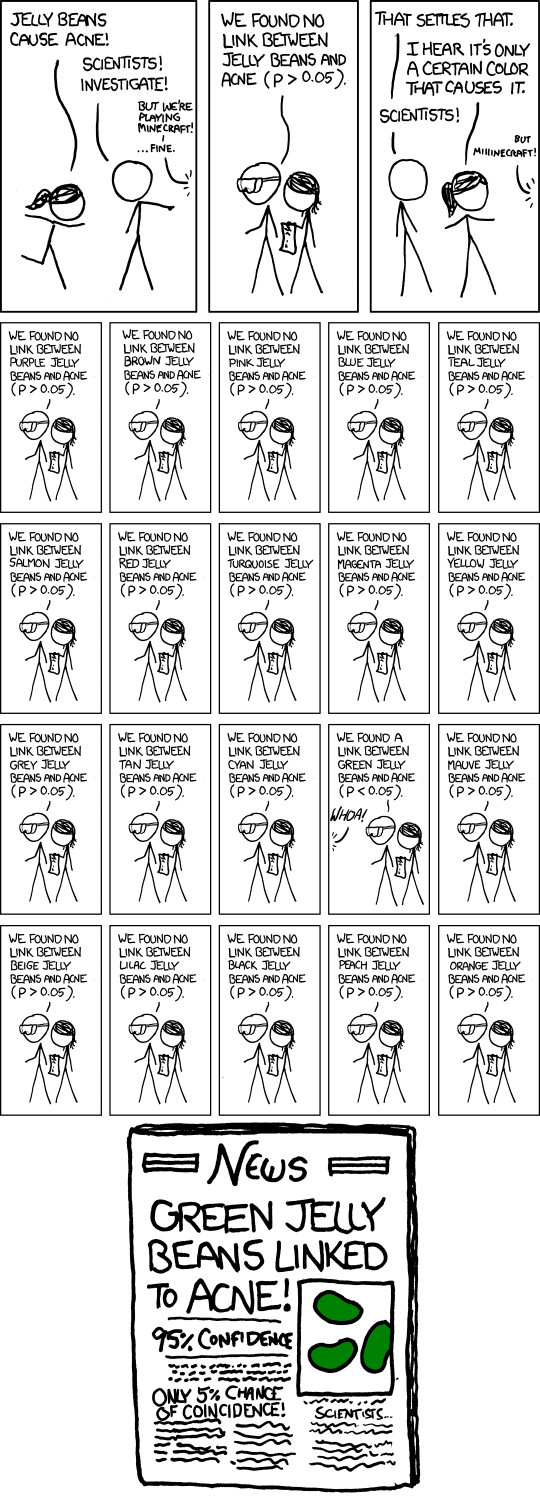
xkcd on p-values
- Significance levels are fairly arbitrary. Sometimes they are used (wrongly) as definitive judgments
- They can even be used to do p-hacking: Searching for “significant” effects in observational data
- In parts of science it has become a “gamed” performance metric.
- The p-value says nothing about effect size!
p-value misinterpretation
p-values do not measure1
- the probability that the studied hypothesis is true
- the probability that the data were produced by random chance alone
- the size of an effect
- the “importance of a result” or “evidence regarding a model or hypothesis” (it is only against the null hypothesis).
Correct:
The p-value is the probability of obtaining test results at least as extreme as the result actually observed, under the assumption that the null hypothesis is correct.
p-values and significance tests, when properly applied and interpreted, increase the rigor of the conclusions drawn from data.2
Classification: Compare logistic regression and decision tree
Purpose of the section
- Go again through the modeling workflow (with
tidymodels) and see that large parts are identical - Look again at the coefficients of a logistic regression model
- Learn the basic idea of a decision tree (you will not learn the details here)
- Do the classification with both models and compare the confusion matrices
Specify recipe and models
For both logistic regression and decision tree:
- We specify a recipe to predict
sexwith all available variables inpenguins- In typical model development, more pre-processing steps are done.
Logistic Regression
peng_logreg specifies to fit with glm (generalized linear model from base R)
Split and fit
For logistic regression and decision tree:
Split into test and training data
Look at fitted logistic regression
# A tibble: 9 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -90.4 16.7 -5.41 0.0000000632
2 speciesChinstrap -6.06 1.90 -3.19 0.00142
3 speciesGentoo -8.65 3.49 -2.48 0.0132
4 islandDream 0.752 1.15 0.656 0.512
5 islandTorgersen 0.380 1.13 0.335 0.737
6 bill_length_mm 0.532 0.150 3.56 0.000378
7 bill_depth_mm 1.79 0.456 3.93 0.0000866
8 flipper_length_mm 0.0552 0.0620 0.891 0.373
9 body_mass_g 0.00693 0.00162 4.27 0.0000193 - What do the categorical predictors tell us? Which are signigficant?
- What do the numerical predictors tell us? Which are signigficant?
- Why is the coefficient for
body_mass_gso small, but highly significant?
Categorical predictors: We have 3 species, 3 island. So, we see 4 new variables, 2 for species and 2 for island (the third is the reference category). Species are significant (p < 0.05), but islands not.
Numerical predictors: flipper_length_mm is insignificant, though its coefficient is larger than for body_mass_g. Reason: values of body_mass_g are larger than those of flipper_length_mm. Body mass differs by much more grams than flipper length differs by millimeters.
What is a decision tree?
══ Workflow [trained] ══════════════════════════════════════════════════════════
Preprocessor: Recipe
Model: decision_tree()
── Preprocessor ────────────────────────────────────────────────────────────────
1 Recipe Step
• step_rm()
── Model ───────────────────────────────────────────────────────────────────────
n=234 (6 observations deleted due to missingness)
node), split, n, loss, yval, (yprob)
* denotes terminal node
1) root 234 117 female (0.50000000 0.50000000)
2) bill_length_mm< 48.15 166 57 female (0.65662651 0.34337349)
4) bill_depth_mm< 18.05 103 14 female (0.86407767 0.13592233)
8) body_mass_g< 4987.5 88 4 female (0.95454545 0.04545455) *
9) body_mass_g>=4987.5 15 5 male (0.33333333 0.66666667) *
5) bill_depth_mm>=18.05 63 20 male (0.31746032 0.68253968)
10) body_mass_g< 3875 29 10 female (0.65517241 0.34482759)
20) flipper_length_mm< 190.5 18 3 female (0.83333333 0.16666667) *
21) flipper_length_mm>=190.5 11 4 male (0.36363636 0.63636364) *
11) body_mass_g>=3875 34 1 male (0.02941176 0.97058824) *
3) bill_length_mm>=48.15 68 8 male (0.11764706 0.88235294)
6) body_mass_g< 5175 35 8 male (0.22857143 0.77142857)
12) bill_depth_mm< 18 9 3 female (0.66666667 0.33333333) *
13) bill_depth_mm>=18 26 2 male (0.07692308 0.92307692) *
7) body_mass_g>=5175 33 0 male (0.00000000 1.00000000) *- A sequence of rules for yes/no decisions
- Selects variables and thresholds which separate the data to predict (here
sex) best - Further details are not the scope of this course
Show rules
We “dig out” the original fitted rpart-object from the workflow-object with peng_tree_fit$fit$fit$fit and plot it:
..y
0.05 when bill_length_mm < 48 & body_mass_g < 4988 & bill_depth_mm < 18
0.17 when bill_length_mm < 48 & body_mass_g < 3875 & bill_depth_mm >= 18 & flipper_length_mm < 191
0.33 when bill_length_mm >= 48 & body_mass_g < 5175 & bill_depth_mm < 18
0.64 when bill_length_mm < 48 & body_mass_g < 3875 & bill_depth_mm >= 18 & flipper_length_mm >= 191
0.67 when bill_length_mm < 48 & body_mass_g >= 4988 & bill_depth_mm < 18
0.92 when bill_length_mm >= 48 & body_mass_g < 5175 & bill_depth_mm >= 18
0.97 when bill_length_mm < 48 & body_mass_g >= 3875 & bill_depth_mm >= 18
1.00 when bill_length_mm >= 48 & body_mass_g >= 5175 - The first three rules would predict
femalefor all observations - The last five rules would predict
malefor all observations
The order of male and female is because sex is a factor with the first level female and the second level male. The probabilities in front of the rule-text are for the second level: male.
Visualize tree
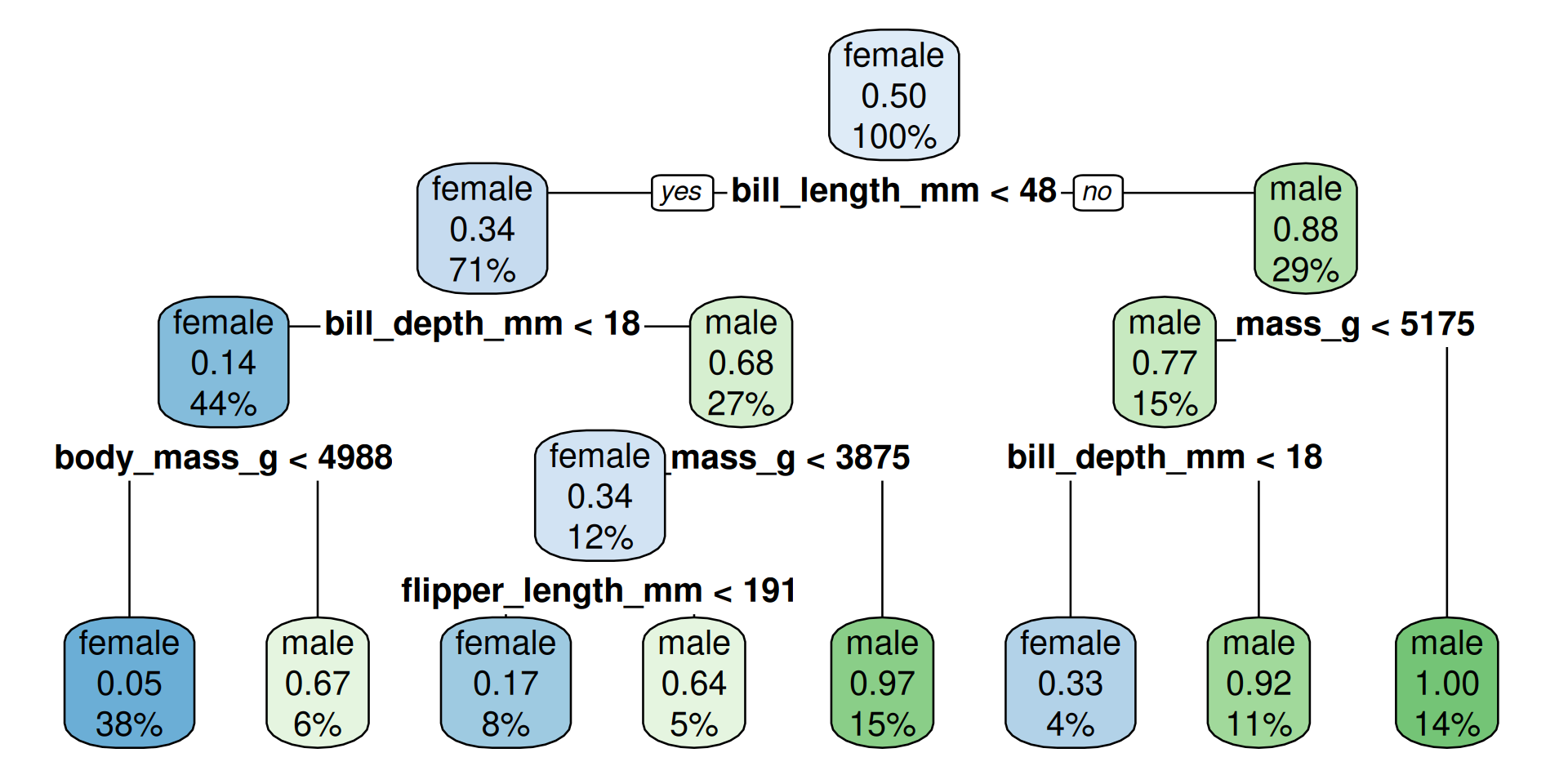
How to read?
- The percentage is the fraction of the total cases in this group
- The probability-number is the fraction of observations which are male in the group
maleoffemaleand color would match the predicted outcome at this decision node
Make predictions for test data
- The logistic regression has more correct predictions.
- Warning: The function
conf_mat(fromyardstickoftidymodels) shows the transposed confusion matrix compared with Wikipedia:Confusion Matrix. Inconf_mat, the true conditions are in columns. The wikipedia convention is that columns are the predicted conditions.
Regression: Compare linear regression and decision tree
Purpose of the section
- Go again through the modeling workflow (with
tidymodels) and see that large parts are identical - Look again at the coefficients of a linear model
- See how the decision tree looks like for a regression problem
- Compare the two most common performance measures for regression models: Root Mean Squared Error (RMSE) and R-squared
Specify recipe and models
- We specify a recipe to predict
body_mass_gwith all available variables inpenguinsand put it in a workflow- Typically, more pre-processing steps are specified here, but we are mostly fine
We can re-use the split and the training and test set.
Fit Regression Models
Look at fitted linear regression
# A tibble: 9 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) -829. 701. -1.18 2.38e- 1
2 speciesChinstrap -268. 109. -2.47 1.44e- 2
3 speciesGentoo 1010. 164. 6.15 3.52e- 9
4 islandDream -14.5 70.0 -0.208 8.36e- 1
5 islandTorgersen -21.4 75.7 -0.283 7.77e- 1
6 bill_length_mm 19.8 8.55 2.32 2.13e- 2
7 bill_depth_mm 53.7 23.9 2.25 2.56e- 2
8 flipper_length_mm 13.6 3.54 3.84 1.58e- 4
9 sexmale 393. 58.5 6.71 1.57e-10- What do the categorical predictors tell us? Which are signigficant?
- What do the numerical predictors tell us? Which are signigficant?
Categorical predictors: We have 3 species, 3 island and 2 sex. So, we see 5 new variables. Species and sex are significant, but islands not.
Numerical predictors: All 3 are significant.
Show rules
..y
3418 when species is Adelie or Chinstrap & sex is female
3961 when species is Adelie or Chinstrap & sex is male
4693 when species is Gentoo & sex is female
5474 when species is Gentoo & sex is male- For each terminal node a certain value is predicted (the mean of the remaining penguins)
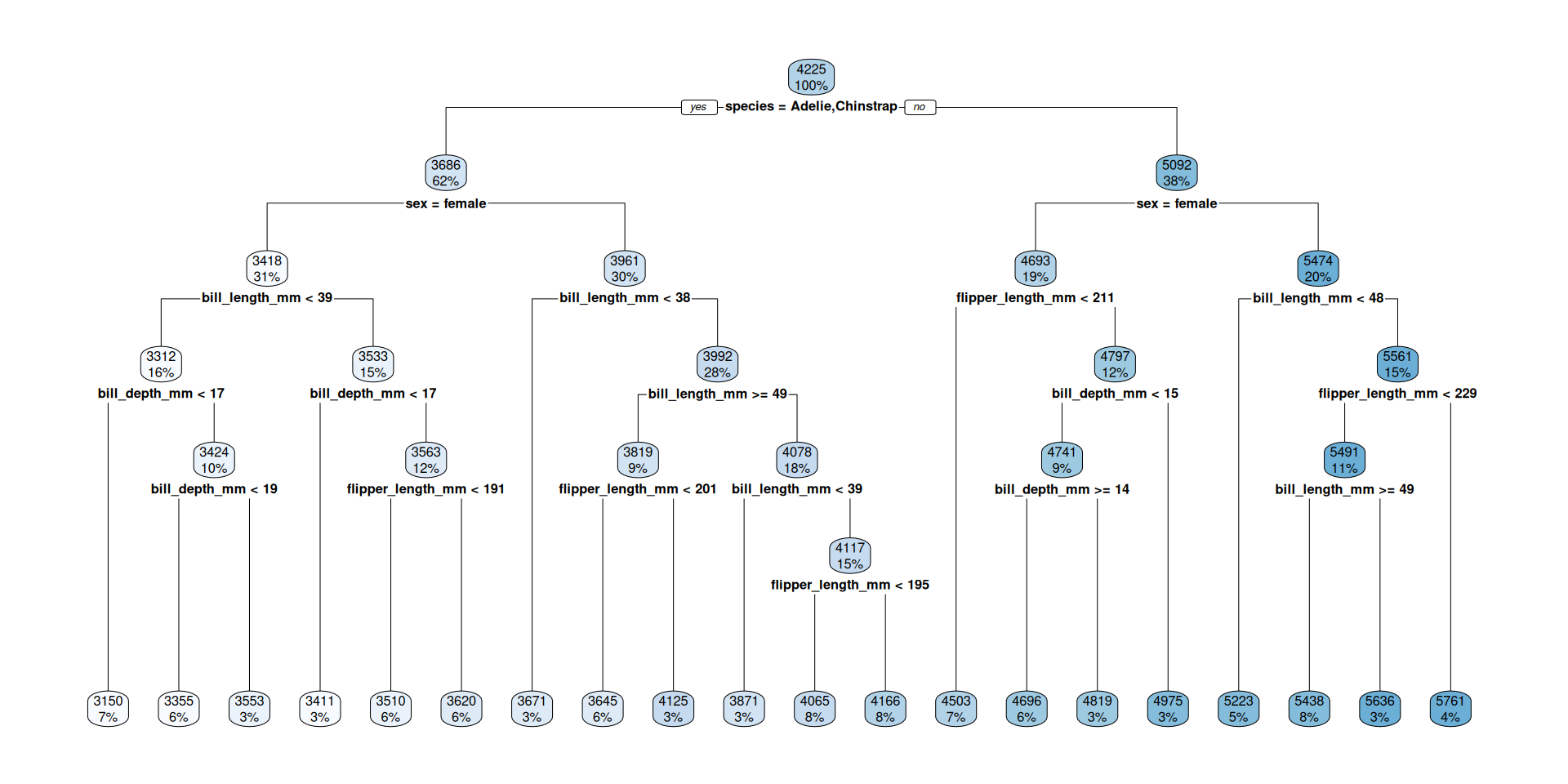
Visualize tree
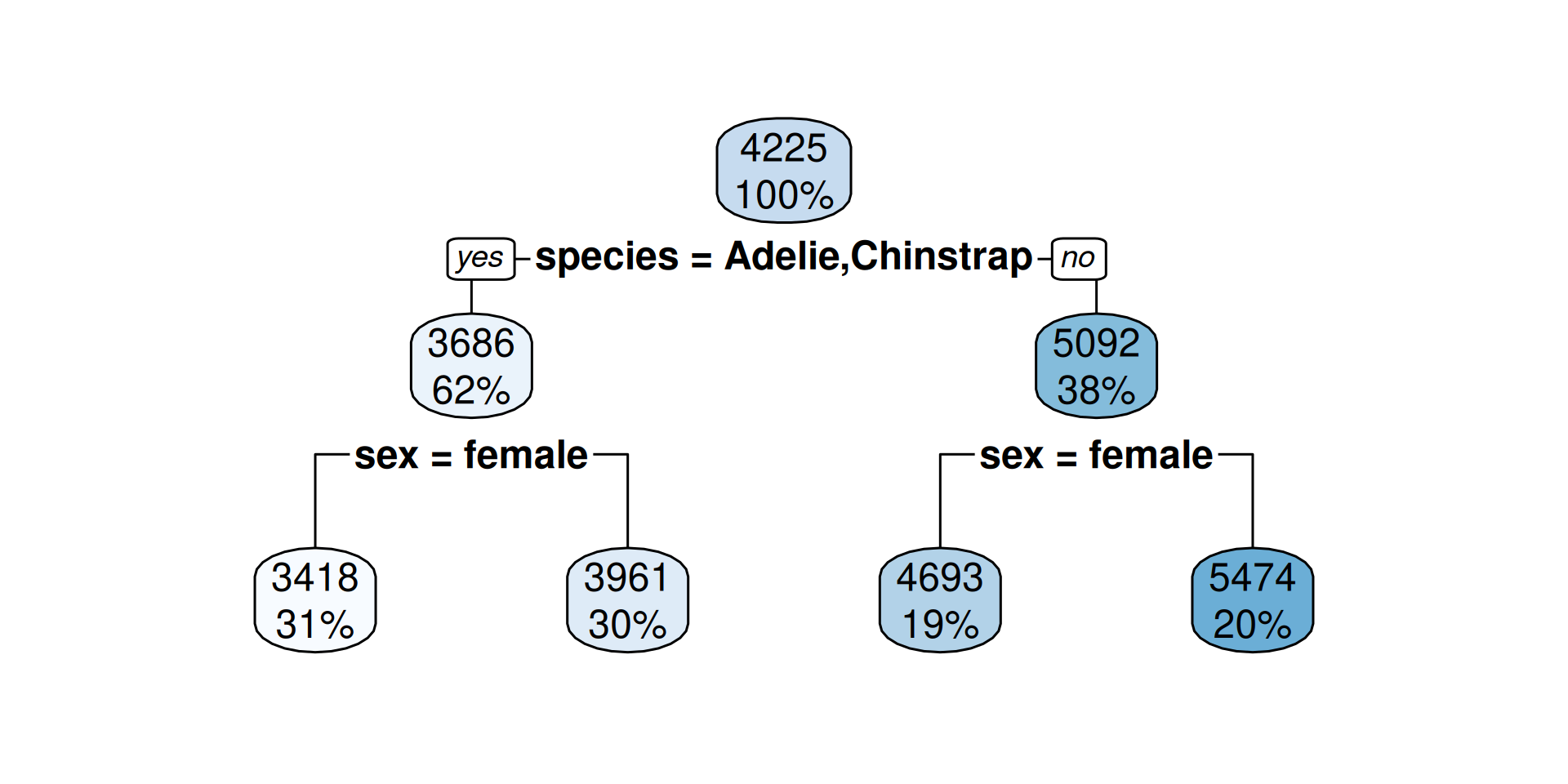
How to read?
- The percentage is the fraction of the total cases in this group
- The number is the predicted outcome (mean body mass) at this decision node
Make predictions for test data
Linear Regression
peng_linreg_pred <-
predict(peng_linreg_fit, peng_test) |>
bind_cols(peng_test)
peng_linreg_pred |>
select(.pred, body_mass_g, everything()) |>
slice(10*(1:10)) # show selected rows# A tibble: 10 × 9
.pred body_mass_g species island bill_length_mm bill_depth_mm
<dbl> <int> <fct> <fct> <dbl> <dbl>
1 3349. 3700 Adelie Torgersen 36.6 17.8
2 3979. 3950 Adelie Dream 38.8 20
3 3902. 3800 Adelie Dream 36.3 19.5
4 4106. 3875 Adelie Torgersen 41.4 18.5
5 3499. 3400 Adelie Dream 40.2 17.1
6 4597. 4150 Gentoo Biscoe 42 13.5
7 4688. 4200 Gentoo Biscoe 45.5 13.9
8 4929. 4850 Gentoo Biscoe 48.5 15
9 4014. 4150 Chinstrap Dream 52 19
10 4108. 4050 Chinstrap Dream 50.7 19.7
# ℹ 3 more variables: flipper_length_mm <int>, sex <fct>, year <int>Decision Tree
peng_regtree_pred <-
predict(peng_regtree_fit, peng_test) |>
bind_cols(peng_test)
peng_regtree_pred |>
select(.pred, body_mass_g, species, sex) |>
slice(10*(1:10)) # show same selected rows# A tibble: 10 × 4
.pred body_mass_g species sex
<dbl> <int> <fct> <fct>
1 3418 3700 Adelie female
2 3961. 3950 Adelie male
3 3961. 3800 Adelie male
4 3961. 3875 Adelie male
5 3418 3400 Adelie female
6 4693. 4150 Gentoo female
7 4693. 4200 Gentoo female
8 4693. 4850 Gentoo female
9 3961. 4150 Chinstrap male
10 3961. 4050 Chinstrap male Regression Performance Evaluation
R-squared: Percentage of variability in body_mass_g explained by the model
Linear Regression
Root Mean Squared Error (RMSE): \(\text{RMSE} = \sqrt{\frac{1}{n}\sum_{i = 1}^n (y_i - \hat{y}_i)^2}\)
where \(\hat{y}_i\) is the predicted value and \(y_i\) the true value. (The name RMSE is descriptive.)
Which model is better in prediction? Linear regression. The R-squared is higher.
What RMSE is better?
Lower. The lower the error, the better the model’s prediction.
It is the other way round than R-squared! Do not confuse them.
Notes:
- The common method to fit a linear model is the ordinary least squares (OLS) method
- That means the fitted parameters should deliver the lowest possible sum of squared errors (SSE) between predicted and observed values.
- Minimizing the sum of squared errors (SSE) is identical to minimizing the mean of squared errors (MSE) because it only adds the factor \(1/n\).
- Minimizing the mean of squared errors (MSE) is identical to minimizing the root mean of squared errors (RMSE) because the square root is strictly monotone function.
Conclusion: RMSE can be seen as a definition of the OLS optimization goal.
Interpreting RMSE
In contrast to R-squared, RMSE can only be interpreted with knowledge about the range and of the response variable! It also has the same unit (grams for body_mass_g).
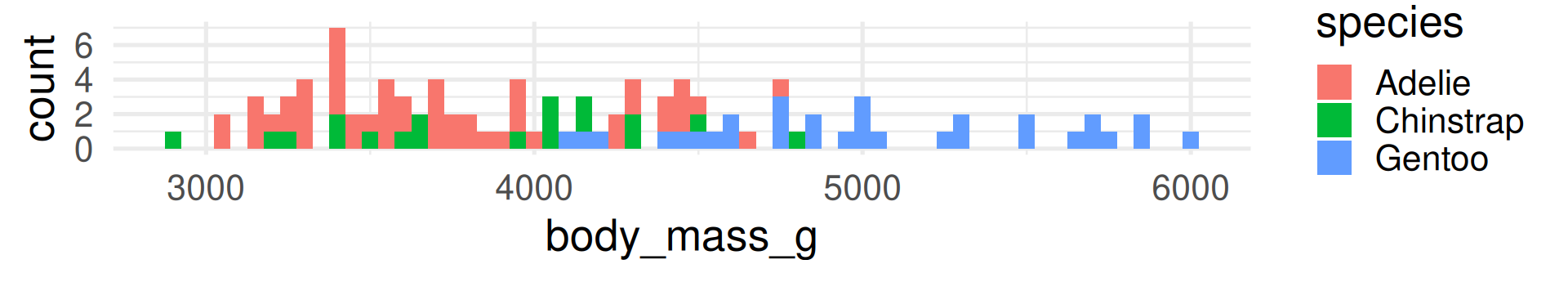
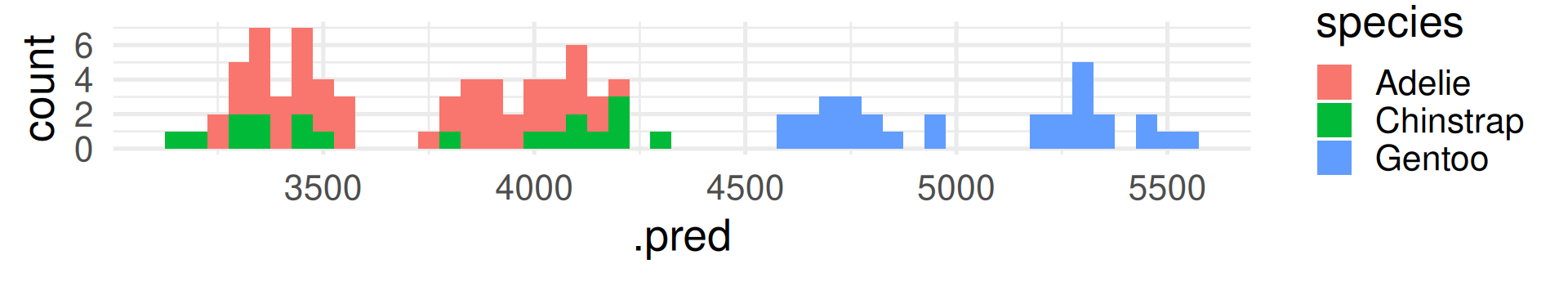
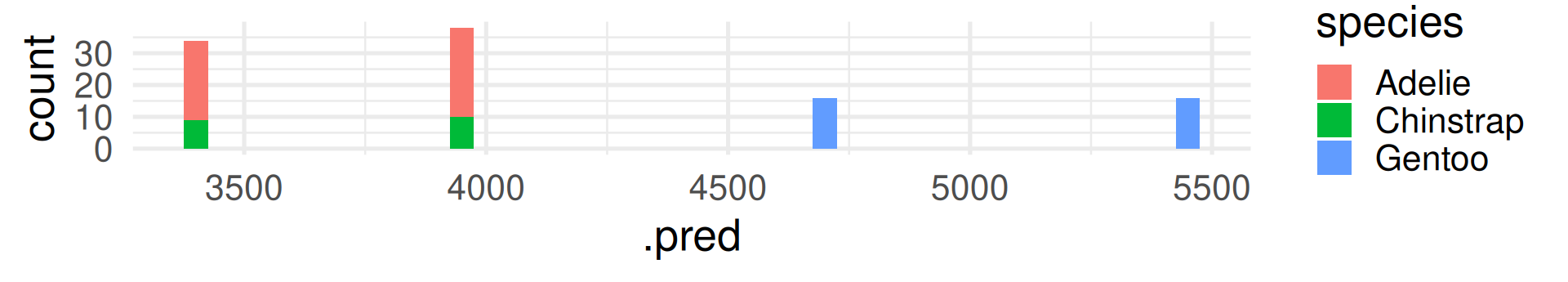
The RMSE shows many grams predicted values deviate from the true value on average. (Taking the squaring of differences and root of the average into account.)
Overfitting with decision trees
Compare training and test data
Linear Regression Predict with test data:
predict(peng_linreg_fit, peng_test) |> bind_cols(peng_test) |>
metrics(truth = body_mass_g, estimate = .pred) |> slice(1:2)# A tibble: 2 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 rmse standard 290.
2 rsq standard 0.861Predict with training data:
Where is the prediction better? (Lower RMSE, higher R-squared)
Performance is better for training data (compare values to testing data of the same model). Why? It was used to fit. The model is optimized to predict the training data.
Make a deeper tree
- In
decision_tree()we can set the maximal depth of the tree to 30.
- The trees we had before were also automatically pruned by sensible defaults.
- By setting the cost complexity parameter to -1 we avoid pruning.1
The deep tree
Compare pruned and deep tree
Pruned decision tree
Predict with training data:
peng_regtree_fit |>
predict(new_data = peng_train) |> bind_cols(peng_train) |>
metrics(truth = body_mass_g, estimate = .pred) |> slice(1:2)# A tibble: 2 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 rmse standard 309.
2 rsq standard 0.857Predict with testing data:
Deep decision tree
Predict with training data:
peng_regtree_deep_fit |>
predict(new_data = peng_train) |> bind_cols(peng_train) |>
metrics(truth = body_mass_g, estimate = .pred) |> slice(1:2)# A tibble: 2 × 3
.metric .estimator .estimate
<chr> <chr> <dbl>
1 rmse standard 247.
2 rsq standard 0.908Predict with testing data:
More in an R-script
Overfitting in spam predicton models.
From decision trees to random forrests.
Move to the script.
What do we mean by “model”? How to be specific
“Model” in Statistical Learning
We already had the difference between variable-based and agent-based models in earlier lectures.
But even in the variable-based model setting of statistical learning, the term model can be more or less abstract:
- Very general: \(Y = f(X_1, \dots, X_m) + \varepsilon\) where \(Y\) is the response variable and \(X_i\) are features which we put in our model: the abstract and unknown function \(f\).
\(\varepsilon\) is the error which we can never explain usually do not know. - More specific: The model \(f\) could already be of a specific type, like logistic regression, a decision tree or other functional forms. As this need not be the real function we may call it assumed model \(\hat f\) For example a linear model \(\hat f(X_1, \dots, X_m) = \beta_0 + \beta_1 X_1 + \dots + \beta_m X_m + \varepsilon\). Now, the model has specified parameters which values are unknown.
More specific: Fitted model
- Fitted model: When we have a data set with values for \(Y, X_1, \dots, X_m\) we can fit values for the parameters \(\hat\beta_0, \dots, \hat\beta_m\) to the data. This is the fitted model \(\hat f\). This is called parameter estimation: We estimate \(\hat\beta_0, \dots, \hat\beta_m\) with the hope that they match the real values \(\beta_0, \dots, \beta_m\) and that the linear model \(\hat f\) matches the real function \(f\).
- Now we could specify further to fit a specific parameterized model with a specific algorithm, and a specific set of hyperparameters, and maybe more …
Sometimes model means only a certain aspect of all these, for example the formula like sex ~ bill_length_mm + bill_depth_mm + flipper_length_mm + body_mass_g + species + island
Take away: “Model” can mean things with very different granularity. That is OK because they are all related and all fit the definition of being a simplified representation of reality.
Be prepared to specify what you mean when you are talking about a model.

MDSSB-DSCO-02: Data Science Concepts